Research
The completion of the Arabidopsis genome in late 2000 revealed that this reference plant coded for over 25,000 genes, the vast majority having no known function. The intervening years has seen a considerable investment by the plant community in attempting to determine the functions and interactions of these encoded proteins. Issues such as protein redundancy and the sheer number of unknown function proteins have been major impediments in attributing function using traditional molecular and biochemical techniques.
|
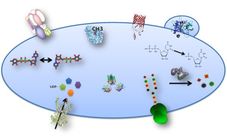
Much of my groups research focuses on the utilization of subcellular proteomic data from compartments involved in plant cell wall biosynthesis (e.g. Golgi apparatus, cytosol and endoplasmic reticulum) to investigate protein function and metabolic partitioning. The investigation of these processes has resulted in the development of novel tools, strategies and basic knowledge that can be used to better understand cell wall biosynthesis in plants.
|
|
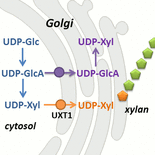
The delivery of nucleotide sugar substrates into the endomembrane for processes such as cell wall biosynthesis and protein glycosylation is critical for plant growth and development. Although plant genomes encode large families of putative nucleotide sugar transporters (NSTs) that deliver the diverse array of nucleotide sugars found in plants, few have been characterized. Recently, we developed a yeast proteoliposome transporter assay coupled to LC-MS, specifically designed to characterize NSTs.
|
|
The Arabidopsis community is currently well-served by online resources, however most have a genetics or genomics emphasis. Over a decade ago, we recognized the need for protein-based resources to assist emerging protein-based technologies, namely subcellular proteomics and the SUBA database. We have now been involved in the establishment of a variety of online tools and portals that often combine public data to enable deeper analyses to occur.
|